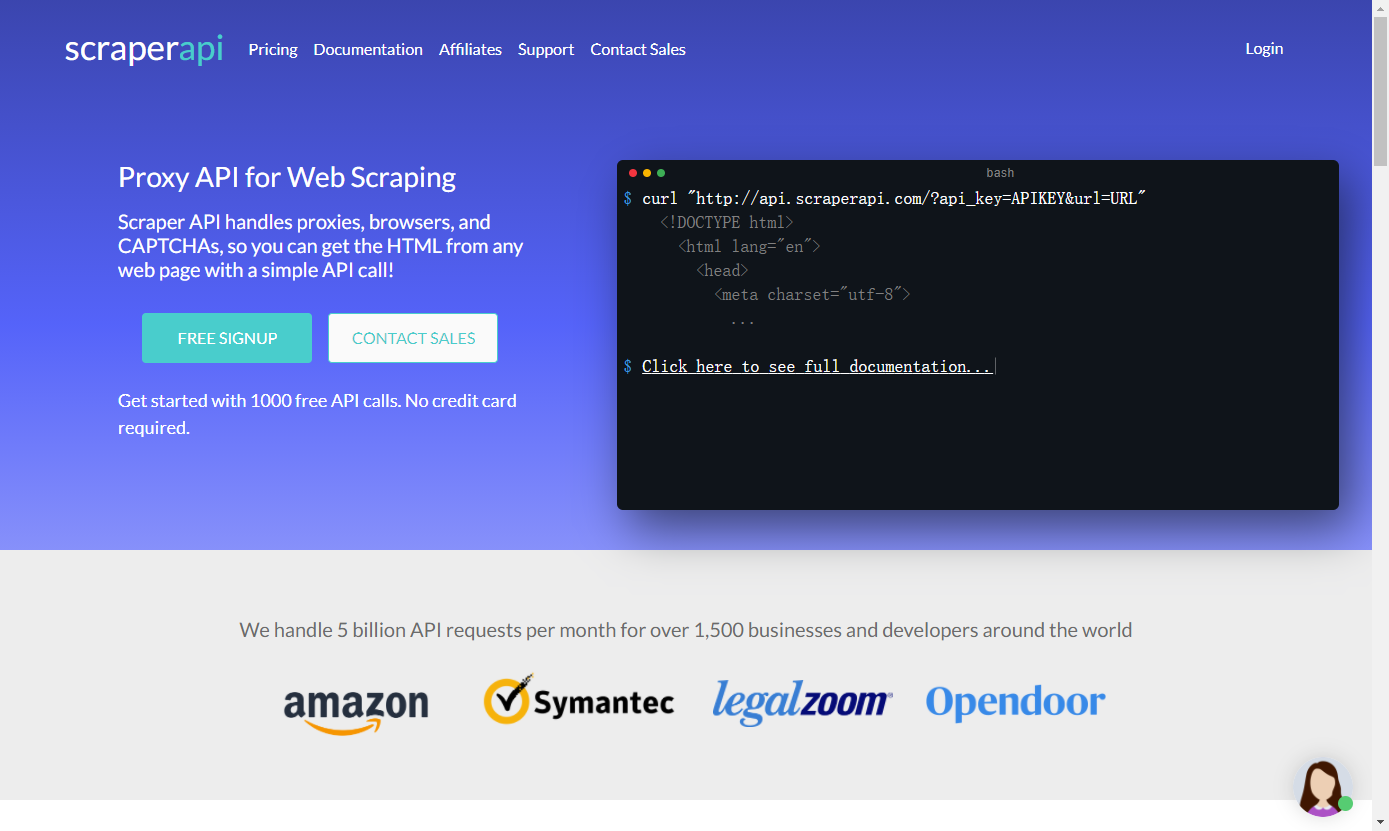
The BRCA2 protein is, through interactions with several other proteins including BRCA1 and PALB2, involved in the repair of DNA double-strand breaks (DSBs) by homologous recombination (Sy et al., 2009). 
Several genetic variants have been shown to increase ∆E3 at the mRNA level (Caputo et al., 2018 Diez et al., 2007), and there is now convincing evidence that near exclusive expression of the ∆E3 transcript (also termed complete exon skipping, or complete ∆E3) is deleterious to protein function and confers high risk of BRCA2-related cancers (Caputo et al., 2018). Skipping of BRCA2 exon 3 (∆E3) is a naturally occurring in-frame splicing event (Peixoto et al., 2009 Thomassen et al., 2012). Interpretation of clinical significance for these spliceogenic variants is difficult as aberrant transcript expression patterns might impact protein function depending on both transcript ratios and the functional integrity of the alternative isoforms. Several naturally occurring alternative splice isoforms of BRCA2 have been reported, and genetic variants have been shown to change the relative expression levels of these alternative transcripts (de Garibay et al., 2014 Fackenthal et al., 2016 Montalban et al., 2018). Although intensive gene testing has been performed for families with suspected hereditary cancer for more than 20 years, there are still considerable challenges for the interpretation of variants with uncertain clinical significance (Spurdle et al., 2012). Pathogenic variants in BRCA2 predispose carriers to breast, ovarian, prostate, and other BRCA2-related cancers (Gayther et al., 1997). 
Our analysis shows the value of applying gene-specific ACMG/AMP guidelines using a points-based approach and highlights the consideration of cryptic splice site usage to appropriately assign PVS1 code strength. Overall, 69/85 (81%) variants were classified using the points-based approach. Cryptic splice site use for acceptor site variants generated a transcript encoding a shorter protein that retains activity. mRNA and mESC analysis combined identified six variants with transcript and/or functional profiles interpreted as loss of function. Variant class was assigned using a gene-specific adaptation of ACMG/AMP guidelines, following a recently proposed points-based system. For each variant, population frequency, bioinformatic predictions, clinical data, and existing mRNA splicing and functional results were collated. 
A mouse embryonic stem cell (mESC) based assay was used to determine the impact of 18 variants on mRNA splicing and protein function. ∆E3 was measured using quantitative analysis. Bioinformatically predicted spliceogenic variants underwent mRNA splicing analysis using minigenes and/or patient samples. This study used multiple evidence types to assess pathogenicity of 85 variants in/near BRCA2 exon 3. It's more flexible than cut when parsing columns of data such as what you're trying to accomplish: $ awk '' data.Skipping of BRCA2 exon 3 (∆E3) is a naturally occurring splicing event, complicating clinical classification of variants that may alter ∆E3 expression. Not all those spaces between the columns look to be tabs, so cut won't be able to do what you want.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
